"Biology has become the most active, the most relevant, and the most personal science, one characterized by extraordinary rigor and predictive power." - John Alexander Moore
AP Biology / College Biology Resources
|
Michelle Obama on the benefit of taking AP classes.
|
What is AP Biology?
Advanced Placement (AP) Biology is a college-level biology course that explores major biological topics in greater depth (evolution, ecology, biochemistry, cell biology, metabolism, genetics, molecular biology, homeostasis, and physiology) while engaging in more advanced laboratory experiences. Why take AP Biology?
|
If you reach a point where you are struggling, please visit the Tips for Taking College Biology Courses page to see if any of that is helpful for you!
Unit 0: Introduction to Biological Science
Experimental Design and Basic Biostatistics
LEARNING TARGETS
- I can design experiments using the scientific method.
- I can represent the data collected from experiments in appropriate and accurate graphs.
- I can analyze the results collected from experiments by formulating scientific claims, evidence, and reasoning.
- I can calculate and interpret the basic descriptive statistics (mean, standard deviation, standard error of the mean) of a data set.
- I can calculate and interpret a Chi-square test to determine potential statistical significance in experimental results.
|
Notes/Reading
|
Key Vocabulary
claim constant control group dependent variable evidence experimental group independent variable negative control positive control reasoning |
|
Key Vocabulary
confidence interval error bars mean sample size/population standard deviation standard error of the mean |
|
Resources - Chi-Square Test
|
Key Vocabulary
Chi-square test critical value degrees of freedom p-value |
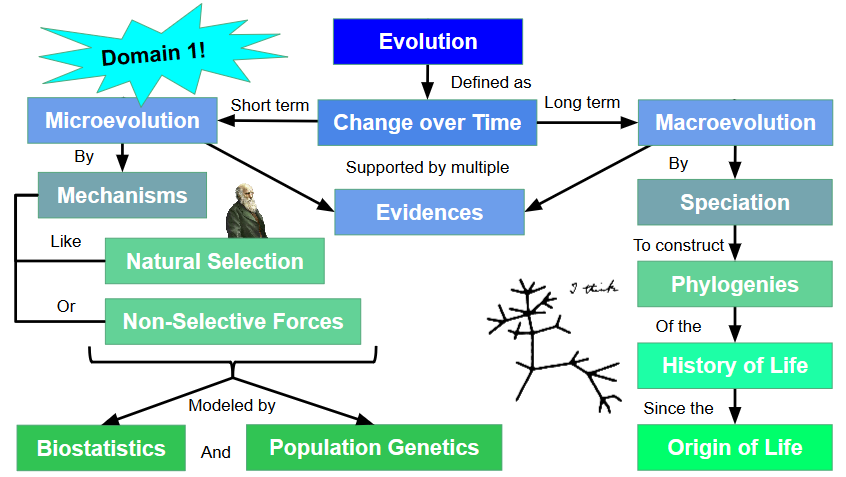
Unit 1: Microevolution
1.1 Evolutionary Mechanisms
LEARNING TARGETS
- I can create a cause-and-effect model of natural selection to relate genotypic diversity, phenotypic variation, environmental pressure, and the differential survival and reproductive success of individuals in populations.
- I can apply descriptive statistics to explain the effect of natural selection or artificial selection on the phenotypic variation in a population.
- I can explain and model how nonselective processes (mutation, genetic drift, gene flow) drive genetic change in a population.
|
Notes/Reading
|
Lab Resources
Study Resources - Natural Selection
Study Resources - Other Mechanisms of Evolution |
Key Vocabulary - Natural Selection
adaptation antibiotic resistance artificial selection coevolution differential survival directional selection disruptive selection evolutionary fitness gene (allelic) frequency genetic variation microevolution mutation natural selection polygenic trait physiological selection predation selection selective pressure stabilizing selection Key Vocab - Other Mechanisms bottleneck effect founder effect gene flow genetic drift sexual dimorphism sexual selection |
1.2 Population Genetics
LEARNING TARGETS
- I can identify and describe the five conditions under which allele and genotype frequencies change in populations.
- I can apply the Hardy-Weinberg model to predict allele and genotype frequencies in a population in genetic equilibrium.
- I can apply a Chi-square test to determine whether a population is in Hardy-Weinberg equilibrium.
|
Notes/Reading
|
Key Vocabulary
dominant vs. recessive gene pool gene vs. allele vs. trait Hardy-Weinberg equilibrium HW variable: 2pq HW variable: p HW variable: p^2 HW variable: q HW variable: q^2 heterozygote advantage homozygous vs. heterozygous locus polymorphism |
Unit 2: Macroevolution
2.1 Speciation
LEARNING TARGETS
- I can apply the biological species concept to identify unique species and critique its limitations.
- I can describe and model the conditions and rates at which speciation can occur under different ecological conditions.
- I can identify and explain prezygotic and postzygotic mechanisms of reproductive isolation that drive speciation.
- I can relate speciation and extinction rates to ecosystem-level and global biodiversity.
|
Notes/Reading
|
Resources
|
Key Vocabulary
adaptive radiation allopatric speciation background extinction rate biological species concept ecological niche ecomorph gradualism hybrid / hybridization hybrid zone macroevolution punctuated equilibrium reproductive isolation ring species species sympatric speciation Types of Barriers
Prezygotic Barriers behavioral isolation ecological isolation gametic isolation geographic isolation mechanical isolation temporal isolation Postzygotic Barriers hybrid breakdown reduced hybrid fertility reduced hybrid viability |
2.2 Phylogeny and Common Ancestry
LEARNING TARGETS
- I can interpret and construct phylogenetic trees based on multiple evidences to infer evolutionary relatedness of organisms.
- I can explain how morphological, biochemical, and geological data provide evidence for organisms' common ancestry.
|
Notes/Reading
|
Resources
Links for Whale Phylogeny Lab
|
Key Vocabulary
analogous structure basal taxon biogeography cladogram comparative embryology convergent evolution divergent evolution homologous structure homoplasy molecular clock monophyletic group (clade) morphology outgroup paraphyletic group parsimony phylogenetics polyphyletic group radiometric (absolute) dating relative dating shared ancestral characteristic shared derived characteristic sister group strata (in geology) transition fossil vestigial structure |
2.3 Origin of Life
LEARNING TARGETS
- I can describe the fundamental molecular and cellular features shared across all domains of life due to universal common ancestry.
- I can describe the scientific evidence that provides support for models of the origin of life on Earth.
|
Notes/Reading
|
Resources - Origin of Life
Resources - Endosymbiosis
Resources - Extinction |
Key Vocabulary
abiogenesis Cambrian explosion chemosynthesis endosymbiotic theory geologic record half-life horizontal gene transfer Last Universal Common Ancestor (LUCA) mass extinction Miller-Urey experiments mitochondrial DNA Oparin-Haldane hypothesis panspermia protocell radiometric dating reducing atmosphere RNA world hypothesis Snowball Earth hypothesis |
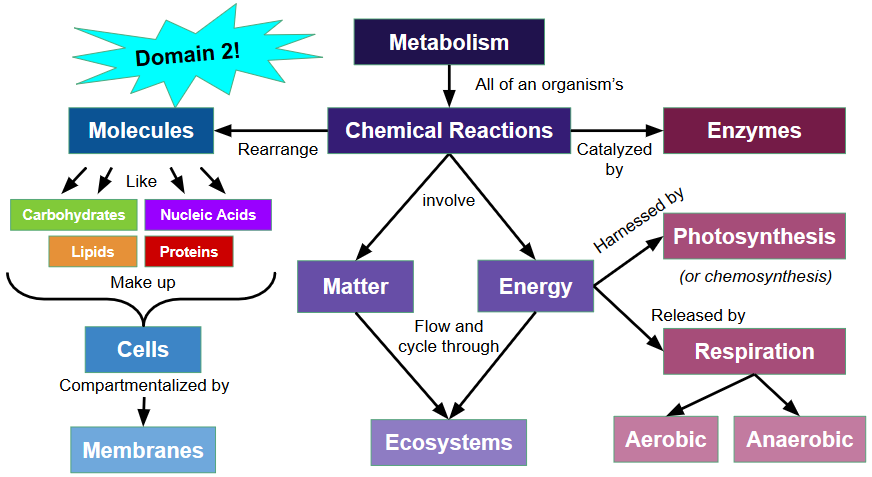
Unit 3: Biochemistry
3.1 Chemical Context of Life and Properties of Water
|
|
|
LEARNING TARGETS
- I can describe how atoms of certain elements are able to combine to compose the structures of organisms.
- I can explain how the properties of water, resulting from water's polarity and hydrogen bonding, affect organisms.
|
Notes/Reading
|
Resources for Chemical Context of Life
Resources for Properties of Water
|
Key Vocabulary - Chemical Context
covalent bond emergent property functional group ionic bond polar vs. nonpolar covalent bond valence electron Key Vocabulary - Water adhesion vs. cohesion buffer capillary action dissociation evaporative cooling hydrogen bond pH = - log [H+] surface tension |
3.2 Biological Molecules
|
|
|
LEARNING TARGETS
- I can describe the properties of monomers and model hydrolysis and dehydration synthesis.
- I can compare and contrast the structures and functions of polymers (carbohydrates, lipids, proteins, nucleic acids) based on the properties of their constitutive monomers.
- I can explain how a change in the subunits of a polymer may lead to changes in structure or function of the macromolecule.
3.3 Metabolism and Enzymes
LEARNING TARGETS
- I can describe the structure and properties of enzymes.
- I can explain and investigate how enzymes function as catalysts to lower the activation energy of a biochemical reaction.
- I can analyze and investigate how changes in the cellular environment can affect enzyme structure and reaction rates.
|
Notes/Reading
|
Resources
Resources - Enzymatic Reaction Rate Labs |
Key Vocabulary
activation energy active site allosteric site anabolism catabolism catalyst coenzyme / cofactor compartmentalization competitive inhibition cooperativity endergonic reaction enzyme-substrate complex exergonic reaction feedback inhibition induced fit model lock-and-key model metabolic pathway metabolism noncompetitive inhibition |
Unit 4: Cells and Membranes
4.1 Cell Structure and Function
LEARNING TARGETS
- I can compare and contrast the compartmentalization of prokaryotic and eukaryotic cells, including the presence or absence of endosymbiotic organelles.
- I can identify and describe the structures, functions, and interrelationships of subcellular components (organelles).
- I can explain how internal membranes and membrane-bound organelles compartmentalize eukaryotic cell activities.
|
Notes/Reading
|
Resources
Resources - Stem Cells and Cell Differentiation
Resources - Endosymbiosis |
Key Vocabulary
cell wall centrosome / centriole chemo-heterotrophic chloroplast cilia vs. flagella cytoskeleton cytosol / cytoplasm endomembrane system endosymbiosis eukaryote (or eukaryotic) extracellular matrix (ECM) Golgi apparatus intercellular junctions lysosome mitochondrion nucleolus nucleus peroxisome photo-autotrophic plasmodesmata plastid prokaryote (or prokaryotic) ribosomes (free and bound) rough endoplasmic reticulum smooth endoplasmic reticulum vacuoles (e.g. central, contractile) vesicles |
4.2 Membrane Structure and Cell Transport
LEARNING TARGETS
- I can describe the molecular structure of the cell membrane and relate these structures to the membrane's selective permeability.
- I can compare and contrast types of cell transport (simple diffusion, facilitated diffusion, osmosis, active transport, bulk transport) based on energy use, concentration gradient, molecule size, molecule polarity, and charge.
- I can explain how the membrane can become polarized, as when Na+/K+ ATPase maintains membrane potential in neurons.
- I can calculate the solute potential of a solution and infer how a cell or organism will osmoregulate in response to the water potential.
4.3 Cell Size
LEARNING TARGETS
- I can calculate the surface area, volume, and surface area-to-volume ratio of a model cell.
- I can explain the effect of surface area-to-volume ratios on the exchange of materials between cells or organisms and their environment.
|
Notes
|
Key Vocabulary
alveoli / lung epithelial cells surface area-to-volume ratio villi / microvilli / intestinal epithelial cells |
Unit 5: Energy
5.1 Free Energy
LEARNING TARGETS
- I can describe the role of free energy in living organisms and explain how living organisms do not violate the first or second laws of thermodynamics.
- I can explain how organisms use coupled reactions, including the ATP cycle, to transform energy between endergonic and exergonic reactions.
|
Notes/Reading
|
Resources for Free Energy
Resources for ATP
Resources for Oxidizing Fuels / Redox Reactions
|
Key Vocabulary
ATP and ADP bioenergetics coupled reaction endergonic reaction enthalpy entropy exergonic reaction first law of thermodynamics Gibbs free energy kinetic energy oxidation potential energy redox reactions (OIL RIG) reduction second law of thermodynamics |
5.2 Cellular Respiration
LEARNING TARGETS
- I can explain how respiration is used by all organisms to use energy stored in biological macromolecules to produce ATP.
- I can explain the role of electron carriers and the electron transport chain in establishing a proton gradient across the inner mitochondrial membrane.
- I can analyze how oxidation during glycolysis, the link reaction, and the Kreb's cycle releases electrons and carbon dioxide.
- I can explain the advantages and disadvantages of organisms using fermentation in anaerobic conditions.
- I can identify, describe, and relate the structures of mitochondria to the process of cellular respiration.
|
Notes/Reading
|
Resources for Aerobic Respiration
Resources for Anaerobic Respiration/Fermentation |
Key Vocabulary
acetyl coenzyme A aerobic respiration alcoholic fermentation anaerobic respiration ATP synthase chemiosmosis chemosynthesis citrate (citric acid) cristae / inner mitochondrial membrane cytochromes decarboxylation dehydrogenase electron carrier electron transport chain electronegativity energy-investment phase energy-payoff phase FAD+ and FADH2 fermentation glycolysis intermembrane space Kreb's cycle / citric acid cycle lactic acid fermentation link reaction mitochondrial matrix NAD+ and NADH oxidative phosphorylation proton gradient pyruvate (pyruvic acid) substrate-level phosphorylation |
5.3 Photosynthesis
LEARNING TARGETS
- I can relate the evolution of photosynthesis in prokaryotes to the oxygenation of the atmosphere and to the evolution of photosynthetic eukaryotic cells.
- I can explain how light-dependent reactions capture energy present in light to generate ATP through an electron transport chain.
- I can explain how captured energy powers the production of carbohydrates from carbon dioxide in the Calvin cycle.
- I can identify, describe, and relate the structures of chloroplasts to the process of photosynthesis.
Unit 6: Cell Division
6.1 Cell Cycle, Mitosis, and Cell Cycle Regulation
LEARNING TARGETS
- I can describe the events of the cell cycle and the role of the cell cycle in growth, healing, and asexual reproduction.
- I can explain how the steps of mitosis work to ensure the passage of a complete set of chromosomes from one generation to the next.
- I can describe regulatory mechanisms for the cell cycle and predict the results of disruptions to the cell cycle.
|
Notes/Reading
|
Resources for Cell Cycle and Mitosis
Resources for Cell Cycle Regulation and Cancer |
Key Vocabulary
anaphase apoptosis binary fission centromere centrosome / centriole / MTOC chromatin cleavage furrow vs. cell plate cytokinesis diploid (2n) G1 phase G2 phase interphase kinetochore metaphase plate mitosis mitotic spindles (microtubules) prophase S phase sister chromatids somatic cell telophase Vocab - Cell Cycle Regulation cancer cell cycle checkpoint cyclin cyclin-dependent kinase density-dependent inhibition metastasis MPF proto-oncogenes tumor suppressor genes |
6.2 Meiosis and Karyotyping
LEARNING TARGETS
- I can explain how the events of meiosis allow for the creation of haploid gametes.
- I can explain how sexual reproduction contributes to genetic diversity via crossing over, independent assortment, and fertilization.
- I can compare and contrast mitosis and meiosis based in terms of chromosome number and type.
- I can explain how certain human disorders can be attributed to specific chromosomal changes like nondisjunction.
|
Notes/Reading
|
Resources for Meiosis and Karyotyping
Resources for Comparing Mitosis and Meiosis
Resources for Chromosomes and Karyotyping |
Key Vocabulary - Meiosis
chiasma crossing over (synapsis) egg / ovum / ovule fertilization gamete germ cell haploid (n) homologous chromosomes independent assortment Meiosis I Meiosis II polar body sister chromatids sperm / spermatozoa spermatogenesis vs. oogenesis tetrad zygote Key Vocabulary - Karyotyping amniocentesis aneuploidy chorionic villi sampling karyotype monosomy / trisomy nondisjunction |
Unit 7: Genetics
7.1 Genetic Inheritance
LEARNING TARGETS
- I can explain the inheritance of genes and traits according to Mendel's laws of segregation and independent assortment, including through monohybrid and dihybrid crosses.
- I can apply the rules of probability to analyze the passage of single-gene traits from parents to offspring.
- I can explain deviations from Mendel's model of inheritance, including incomplete dominance, codominance, sex linkage, multiple alleles, polygenic traits, phenotypic plasticity, and non-nuclear inheritance.
- I can predict the pattern of inheritance from data, including pedigrees.
- I can explain how certain human genetic disorders can be attributed to the inheritance of a single affected/mutated allele.
7.2 Chromosomal Inheritance
LEARNING TARGETS
- I can explain how the chromosomal basis of inheritance provides an understanding of the pattern of transmission of genes from parents to offspring.
- I can explain how linked genes deviate from Mendel's laws and predicted ratios.
- I can predict the pattern of inheritance (sex-linked and genetically linked genes) from data.
|
Notes/Reading
|
Resources
|
Key Vocabulary
aneuploidy barr body chromosomal mutations gene mapping hemizygous histone linked genes locus (of a gene) monosomy / trisomy mutant vs. wild type nondisjunction nucleosome parental types polyploidy recombinant types recombination frequency sex-linked traits Thomas Hunt Morgan x inactivation |
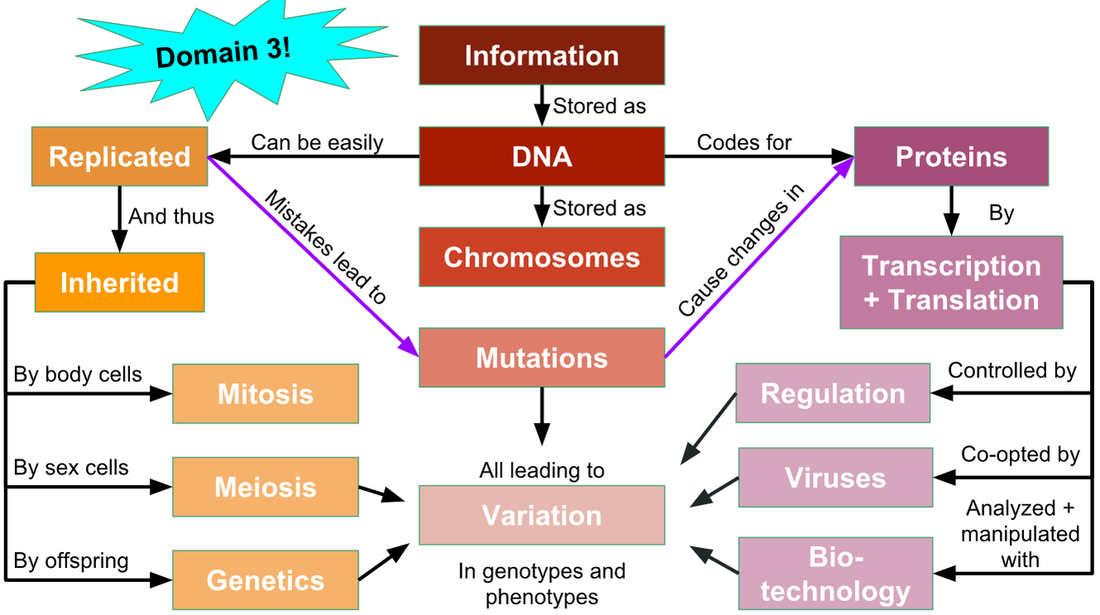
Unit 8: Gene Expression
8.1 DNA Structure, Function, and Replication
LEARNING TARGETS
- I can describe the structure of DNA, including nucleotide structure, nucleotide variety, directionality, and bonding.
- I can use experimental evidence to support the claim that DNA is the molecule containing heritable information.
- I can describe the mechanisms of DNA replication that allow genetic information to be copied for transmission between generations.
|
Notes/Reading
|
Key Vocabulary - DNA Structure
3' (three prime) vs. 5' (five prime) antiparallel Avery, McCarty, MacLeod bacteriophage Chargaff's Rule Frederick Griffith Hershey and Chase hydrogen bond James Watson and Francis Crick nucleotides (the 4 kinds and their 3 parts) phosphodiester bond purines vs. pyrimidines Rosalind Franklin transformation factor |
|
Key Vocabulary - DNA Replication
bidirectional DNA polymerase I DNA polymerase III helicase leading vs. lagging strands ligase Meselson-Stahl experiment Okazaki fragments origin of replication primase and RNA primers replication bubble replication fork semiconservative model SSB topoisomerase |
8.2 Gene Expression
LEARNING TARGETS
- I can describe the mechanisms by which genetic information flows from DNA to mRNA through transcription.
- I can describe post-transcriptional modifications made to mRNA in eukaryotes.
- I can describe the mechanisms by which genetic information flows from mRNA to a protein through translation.
- I can describe types of mutations and explain how these changes in the genotype may or may not result in changes in phenotypes.
|
Notes/Reading
|
Resources for Protein Synthesis
|
Key Vocabulary - Basics
Beadle and Tatum's experiment central dogma gene expression / protein synthesis Key Vocabulary - Transcription antisense strand / template strand mRNA promoter sequence and TATA box RNA polymerase sense strand / coding strand terminator sequence transcription (and 3 steps) transcription factors Key Vocabulary - RNA Processing capping / 5' cap exon intron polyadenylation / poly-A tail ribozyme RNA processing RNA splicing and alternative splicing spliceosome / snRPS Key Vocabulary - Translation anticodon codon polypeptide polyribosome protein processing ribosome, rRNA, ribosomal subunits ribosomes (free and bound) start codon and initiator tRNA stop codon and release factor (RF) translation (and 3 steps) tRNA |
|
Key Vocabulary - Mutations
frameshift mutation missense mutation mutagen nonsense mutation point mutation silent mutation somatic vs. germline mutation |
8.3 Regulating Gene Expression
LEARNING TARGETS
- I can explain the roles of operons in controlling prokaryotic gene expression.
- I can explain the roles of epigenetic changes (histone acetylation and DNA methylation), transcription factors, and microRNA molecules in eukaryotic gene expression.
- I can relate differential gene expression to cell differentiation in multicellular organisms.
|
Notes/Reading
|
Resources - Gene Regulation (General)
Resources - Prokaryotic Gene Regulation Resources - Eukaryotic Gene Regulation Resources - Development
Resources - Stem Cells and Cell Differentiation |
Key Vocabulary - Prokaryotes
corepressor inducer inducible gene (lac operon) negative regulation operator operon / operon model positive regulation promoter repressible gene (trp operon) repressor Key Vocabulary - Eukaryotes acetylation activator alternative RNA splicing differential gene expression euchromatin gene silencing heterochromatin histone methylation microRNA (miRNA) nucleosome protease / proteosome RNA interference (RNAi) transcription factor ubiquitin Key Vocabulary - Cell Differentiation
apoptosis bicoid blastulation / blastula body axes cleavage concentration gradients cytoplasmic determinants differential gene expression ectoderm vs. endoderm vs. mesoderm evo-devo gastrulation / gastrula Hox genes (homeotic genes) induction master regulatory genes totipotent / pluripotent / multipotent zygote cell differentiation transcription factors |
8.4 Viruses and Gene Expression
LEARNING TARGETS
- I can explain how retroviruses have an alternate flow of genetic information (from RNA to DNA) by using reverse transcriptase.
- I can explain how viral DNA integrates into host genomes and becomes transcribed and translated for the assembly of viral progeny.
|
Notes/Reading
|
Resources
|
Key Vocabulary
bacteriophage capsid / capsomeres horizontal gene transfer lysogenic cycle lytic cycle HIV membrane envelope pathogenicity / virulence prophage provirus retrovirus reverse transcriptase temperate phage transduction transformation / transfection virus vaccine |
8.5 Biotechnology
LEARNING TARGETS
- I can... explain how restriction enzymes are used to digest DNA samples into segments with predictable endings.
- I can... explain how polymerase chain reaction (PCR) is used to amplify segments of DNA.
- I can... explain how gel electrophoresis can be used to separate DNA fragments for the purpose of comparison.
- I can... interpret DNA profile/fingerprint results given a specific context or scenario.
- I can... explain the procedures and uses for making recombinant DNA and transforming bacteria.
- I can... explain the procedures and uses for DNA sequencing.
|
Notes/Reading
|
Key Vocabulary - DNA Profiling
DNA profiling / DNA fingerprinting gel electrophoresis restriction enzyme RFLP Southern blotting Western blotting Key Vocabulary - PCR PCR (polymerase chain reaction) Key Vocabulary - Genetic Engineering bacterial transformation cDNA library cloning vector gene cloning gene therapy genomic library plasmid restriction enzyme sticky vs. blunt ends Key Vocabulary - DNA Sequencing DNA barcoding DNA probe Next-Generation sequencing nucleic acid hybridization Sanger sequencing |
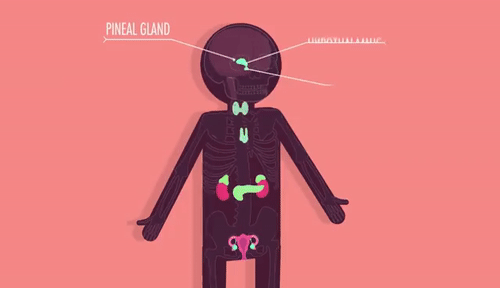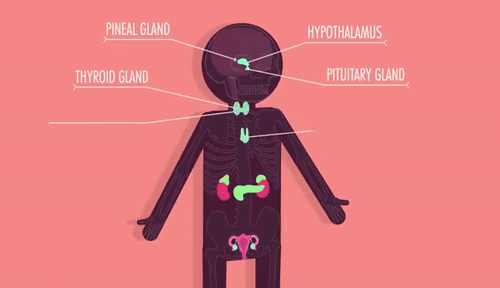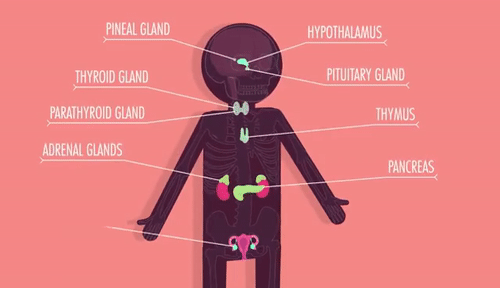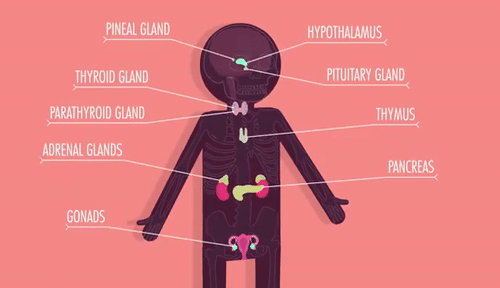
Unit 9: Cell Communication and Homeostasis
9.1 Cell Communication
(LEARNING TARGETS)
|
Notes/Reading
|
Key Vocabulary
adenyl cyclase antigen-presenting cell autocrine biofilm chemotaxis cyclic AMP endocrine G-protein linked receptor gap junction hydrophilic vs. hydrophobic inositol triphosphate juxtacrine kinase ligand ligand-gated ion channels neurotransmitter paracrine phospholipase C phosphorylation and dephosphorylation phosphorylation cascade phototaxis plasmodesmata protein kinase A (PKA) quorum sensing receptor receptor tyrosine kinase second messenger signal amplification signal transduction synapse |
9.2 Homeostasis and Feedback Loops
(LEARNING TARGETS)
|
Notes/Reading
|
Resources
|
Key Vocabulary
control center effector homeostasis negative feedback normal range positive feedback receptor response set point stimulus variable |
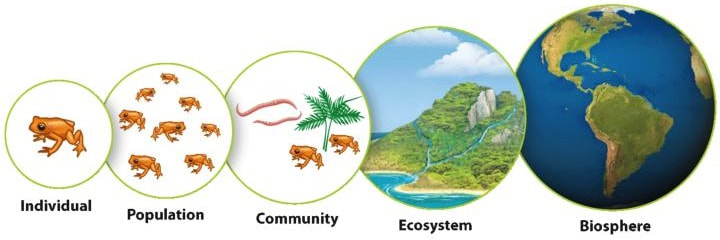
Unit 10: Ecology
10.1 Behavior and Responses
LEARNING TARGETS
- I can explain how organisms respond to changes in their environment through behavioral and physiological mechanisms.
- I can explain how behavioral responses affect an organism's overall fitness and may contribute to the success of the population.
|
Notes/Reading
|
Key Vocabulary
acclimitization adaptation behavior chemotaxis circadian rhythm conditioning dominance hierarchy estivation gravitropism (geotropism) habituation hibernation hydrotropism imprinting innate behavior learned behavior mating ritual migration pheromones photoperiodism phototaxis phototropism proximate explanation stimulus taxis territorial behavior thigmotropism torpor ultimate explanation waggle dance |
10.2 Population Ecology
(LEARNING TARGETS)
|
Notes/Reading
|
Resources
|
Key Vocabulary
abundance (N) age structure diagram biotic potential carrying capacity crude birth rate crude death rate density-dependent limiting factors density-independent limiting factors dynamic equilibrium exponential growth growth rate J-curve K-selected lag phase logistic growth population population density population distribution r-selected replacement level S-curve sex ratio survivorship curves |
10.3 Community Ecology and Changes to Communities
LEARNING TARGETS
- I can explain how species interactions (competition, predation, and symbiosis) influence community structure.
|
Notes/Reading
|
Resources
Resources
|
Key Vocabulary
altruism amensalism commensalism community competitive exclusion fundamental niche generalist species herbivory interspecific competition intraspecific competition mutualism niche niche partitioning optimal range parasitism predation predator-prey curve realized niche specialist species symbiosis tolerance range |
10.4 Energy Flow through Ecosystems
LEARNING TARGETS
- I can describe strategies organisms use to acquire and use energy, regulate body temperature, regulate metabolism based on temperature and body size, and gain or lose mass based on energy availability.
- I can compare how the activities of photosynthetic autotrophs, chemosynthetic autotrophs, and heterotrophs enable the flow of energy within an ecosystem.
- I can interpret food chains and food webs to predict how changes in energy availability affect populations and ecosystems.
|
Notes/Reading
|
Resources
|
Key Vocabulary
10% rule autotroph/producer biomass carnivore cellular respiration chemosynthesis decomposer detritivore ectotherm endotherm entropy First Law of Thermodynamics food chain and food web gross primary production herbivore heterotroph/consumer kinetic energy metabolic rate metabolism net primary production omnivore photosynthesis potential energy primary production Second Law of Thermodynamics secondary production trophic cascade trophic pyramid |
10.5 Biodiversity
LEARNING TARGETS
- I can describe and measure the species composition and species diversity of a community, including using the Simpson's diversity index.